- needhelp@info.com
- 88 broklyn silver street, USA
15Dec2025
-
15Dec2025
Beyond the Lab: Federated, Interpretable Trees That Don’t Leak Your Data
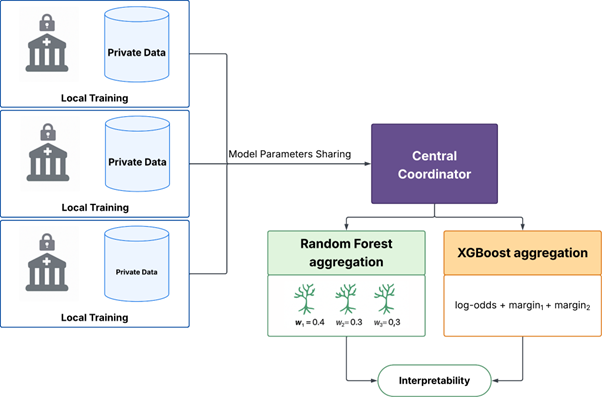
A privacy-aware framework for Random Forests & XGBoost in distributed settings, built for practitioners who need auditability, not black boxes.
Most federated learning toolkits focus on deep nets and often overlook interpretability. If you work with regulated data (e.g. healthcare or finance), that’s a non-starter. This work delivers a federated, PySyft-based approach for tree models such as Random Forests and XGBoost, that keeps data on site, aggregates models sanely and preserves explanations that compliance teams can actually read.
🔍 The Challenge
- Data can’t move. GDPR/HIPAA, institutional trust and real governance barriers kill centralization.
- Tree-based models are under-served. Most FL stacks target gradients; trees need different aggregation and explanation tooling.
- Interpretability matters. You need feature importances and decision paths that survive federation, not glossy dashboards.
🛠️ Our Solution
We built a federated framework that lets hospitals train Random Forest and XGBoost models together without sharing patient data. Each hospital trains locally, then sends only model components—never raw records—to a coordinator. The coordinator combines these intelligently: sampling trees proportionally for Random Forests or averaging prediction scores for XGBoost in ways that preserve mathematical consistency.
This tool computes feature importance rankings entirely within each hospital, aggregating only summary statistics. Clinicians get transparent explanations showing which factors drive predictions, while patient data stays local.
Authored by: Alexandre Cotorobai
In collaboration with: Jorge Miguel Silva, José Luís Oliveira
Supported by: IEETA (Unit 00127), FCT; EC grant 101081813 (Genomic Data Infrastructure).
#FederatedLearning #InterpretableML #Privacy #TreeModels #MLEngineering
